Introduction to R
(Part I)
Giuseppe Arena
Tilburg University
2024-01-11
Who am I?
- Giuseppe Arena, work as Researcher at the Department of Methodology and Statistics (TSB)
- Passionate about statistics and programming
- Background studies: statistics, biostatistics and social network analysis
- Creator and maintainer of several R packages

Goals of this workshop
![]()
![]() understanding
understanding
basic R language
![]() defining
defining
data structures
![]()
writing
functions
![]()
![]()
exploring
data
![]()
visualizing
data
![]()
running
statistical analyses

 understanding
understandingbasic R language
 defining
definingdata structures

writing
functions


exploring
data
visualizing
data
running
statistical analyses
What is R?
Programming language, dialect of the S language, designed in the 90s by Robert Gentleman and Ross Ihaka
Framework suitable for statistical modeling , data processing, analysis, and visualization
Programming language, dialect of the S language, designed in the 90s by Robert Gentleman and Ross Ihaka
Framework suitable for statistical modeling , data processing, analysis, and visualization
Why R?
- free
- open
- simple
- interactive
- versatile
From simple to more complex analyses
Calculating the mean
mean(c(1,2.3,5,3.2,5.4,3,2.1,7,4.8,2))
Multilevel linear regression
lme4::glmer(income ~ (1|family) + condition, data = people)
mean(c(1,2.3,5,3.2,5.4,3,2.1,7,4.8,2))
lme4::glmer(income ~ (1|family) + condition, data = people)
R is widely used
![]() Academic research
Academic research
![]() Social Sciences
Social Sciences
![]() Finance and Economics
Finance and Economics
![]() Bioinformatics
Bioinformatics
 Academic research
Academic research
 Social Sciences
Social Sciences
 Finance and Economics
Finance and Economics
 Bioinformatics
Bioinformatics

Getting Started with R
R GUI and RStudio

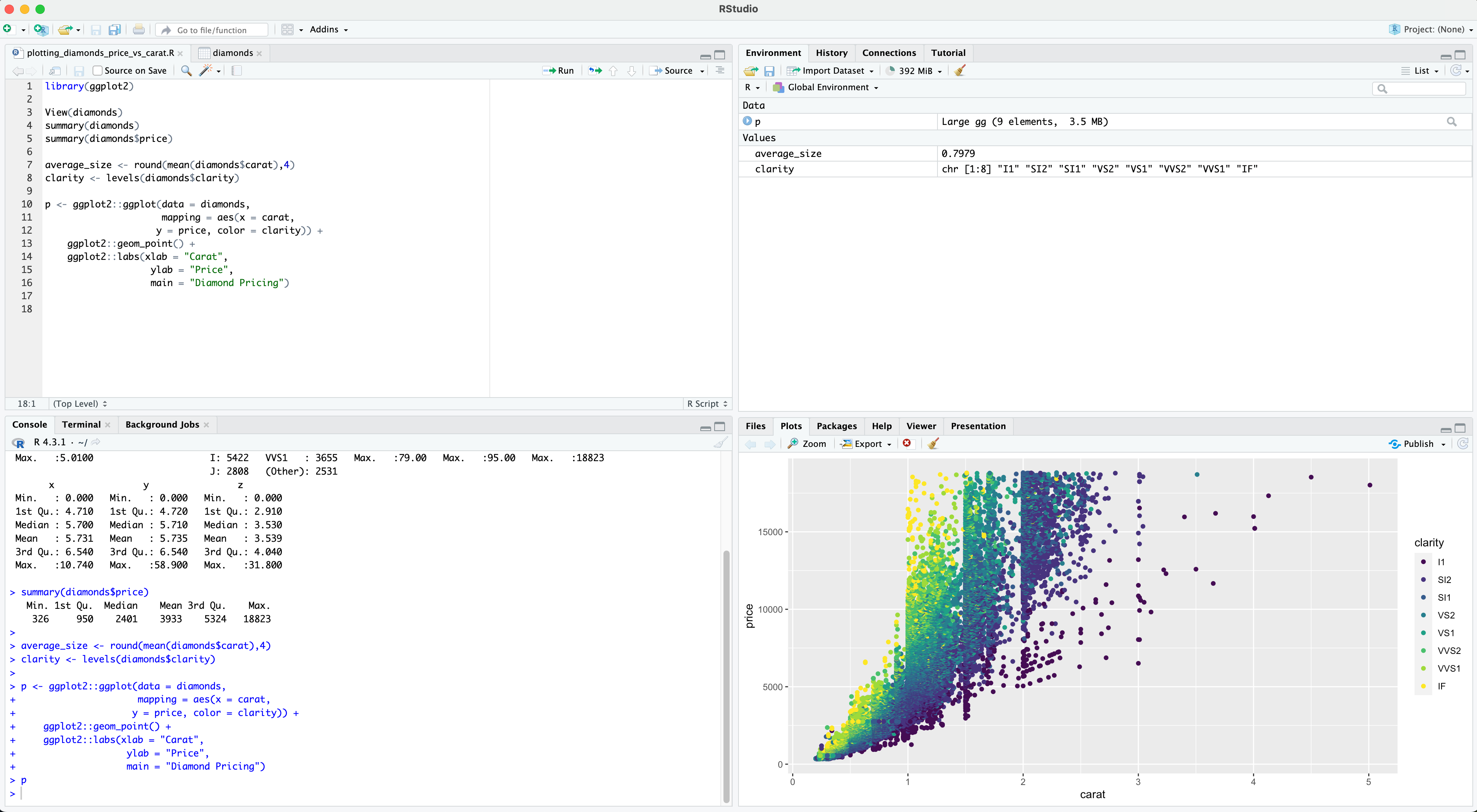
RStudio User Interface

Source code (syntax)
Interactive console
Environment (variables)
Graphics (plots)
R console...
... is interactive!
> "Luke"
[1] "Luke"
> 3.1416
[1] 3.1416
> 3 + 2
[1] 5
> # I am a comment (start a line with # to write any comment)
> I am not a comment
Error: unexpected symbol in "I am"
R Packages
R packages are expansions packs for R and include:
![]()
functions
adding specific functionalities
![]() documentation describing how to use the functions
documentation describing how to use the functions
![]()
data
used for the examples
R comes with base packages: base, utils, datasets, graphics, grDevices, and others
R packages are expansions packs for R and include:

functions
adding specific functionalities
 documentation describing how to use the functions
documentation describing how to use the functions

data
used for the examples
R comes with base packages: base, utils, datasets, graphics, grDevices, and others
Data Structures and Functions
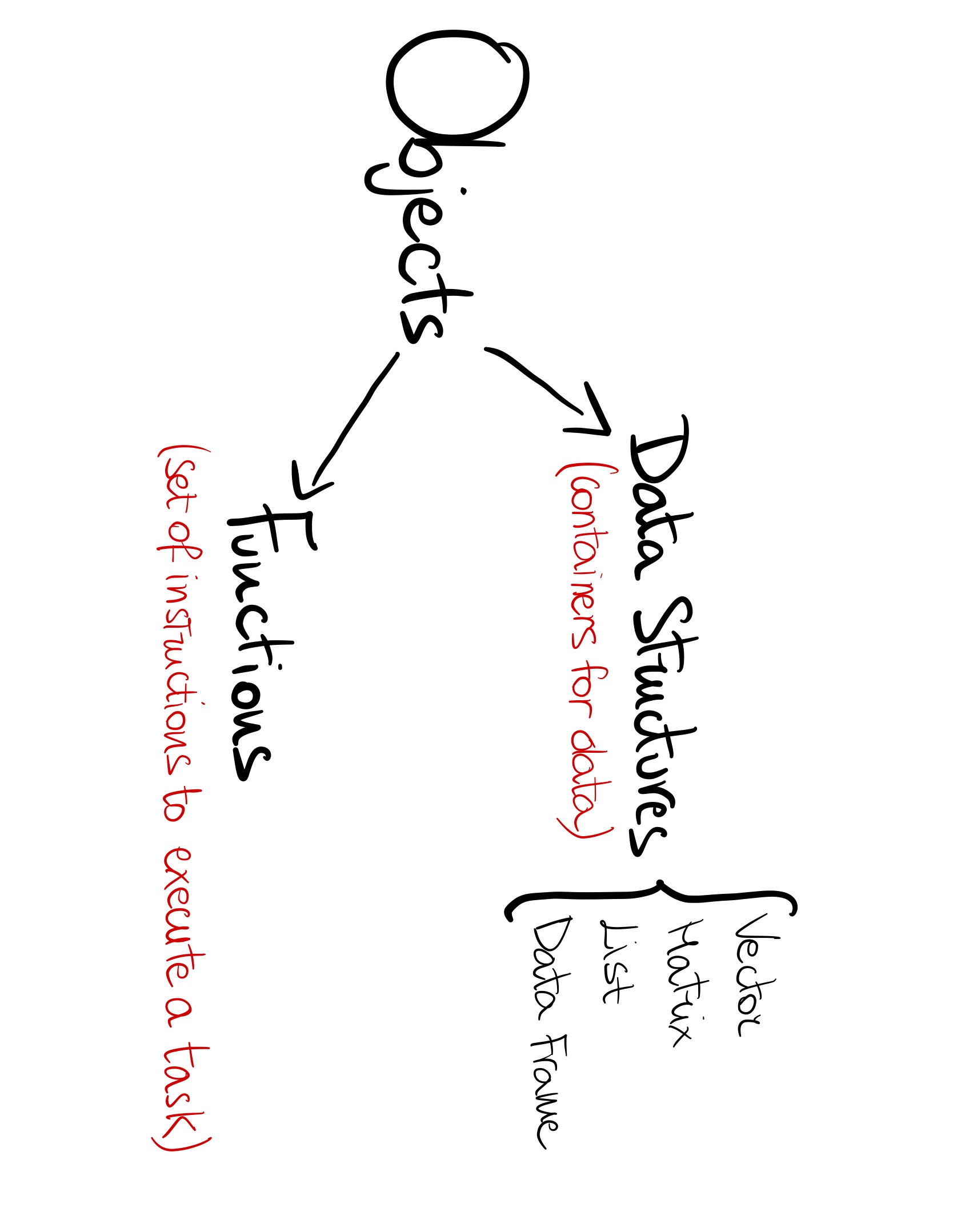
Data Structures
Data Type
A data type specifies how the data is stored in memory and what operations can be performed with that data. Each type has its own characteristics and uses.
Numeric
> 3.4
[1] 3.4
Character
> "workshop"
[1] "workshop"
Integer
> 15L
[1] 15
Logical
> TRUE
[1] TRUE
> FALSE
[1] FALSE
Vector
A vector is a concatenated sequence of elements. Elements in a vector must be of the same data type (numeric, character, logical, etc.). You can create a vector using the c( ) function and separate elements using a comma.
Numeric vector
> c(1.0, 2.3, 0.4, 3.1, 5.1, 4.2, 1.8)
[1] 1.0 2.3 0.4 3.1 5.1 4.2 1.8
Character vector
> c("Luke","Mark","Richard","Hannah","Paul")
[1] "Luke" "Mark" "Richard" "Hannah" "Paul"
Matrix
A matrix is a two-dimensional data structure with rows and columns. All elements in a matrix must be of the same data type. You can create a matrix using the matrix( ) function.
Numeric matrix
> matrix(data = c(1.0, 1.33, 1.66, 2.0, 2.33, 2.66,
3.0, 3.33, 3.66), nrow = 3, ncol = 3)
[,1] [,2] [,3]
[1,] 1.00 2.00 3.00
[2,] 1.33 2.33 3.33
[3,] 1.66 2.66 3.66
The correlation matrix is an example of numeric matrix.
List
A list is a data structure that can contain elements of different data types and structures. Each element in a list can be a vector, matrix, data frame, or another list. You can create a list using the list( ) function.
Example of list
> list(v1 = c(1.1, 2.1, 3.1),
v2 = c("Luke", "Mark"),
m1 = matrix(c(1.1, 2.1, 3.1, 4.1), nrow = 2, ncol = 2))
$v1
[1] 1.1 2.1 3.1
$v2
[1] "Luke" "Mark"
$m1
[,1] [,2]
[1,] 1.1 3.1
[2,] 2.1 4.1
Data frame
A data frame is a tabular data structure with rows and columns, similar to a spreadsheet and commonly used for storing datasets. Columns are named and can have different data types. You can create a data frame using the data.frame( ) function.
Example of data frame
> data.frame(name = c("Luke", "Mark", "Hannah"),
age = c(22, 30, 29),
score = c(8.5, 7.5, 9.2))
name age score
1 Luke 22 8.5
2 Mark 30 7.5
3 Hannah 29 9.2
The vectors defining the columns must have the same length.
Factor
a factor is used to represent a categorical variable in statistical modeling and analysis. A factor has levels that indicate the values that the categorical variable can assume and it is usually created from a character vector. You can create a factor using the factor( ) function.
Example of factor
> # eye color - character vector
> c("brown", "blue", "brown",
"brown", "green", "green", "brown")
[1] "brown" "blue" "brown" "brown" "green" "green" "brown"
> # eye color - factor
> factor(c("brown", "blue", "brown",
"brown", "green", "green", "brown"))
[1] brown blue brown brown green green brown
Levels: blue brown green
Variables
Variables
A variable is a symbolic name given to an object created in R. Variables are used to store and manipulate data and are created by assigning a name to a particular value or object using the assignment operator '
Creating variables name, score and age
> name <- c("Luke","Mark", "Hannah")
> name
[1] "Luke" "Mark" "Hannah"
> score <- c(8.5, 7.5, 9.2)
> score
[1] 8.5 7.5 9.2
> age <- c(22, 30, 29)
> age
[1] 22 30 29
Creating data frame sample_df
> sample_df <- data.frame(name = name,
age = age,
score = score)
> sample_df
name age score
1 Luke 22 8.5
2 Mark 30 7.5
3 Hannah 29 9.2
You can also assign an object to a variable by using the operator '='.
name = c("Luke","Mark", "Hannah")
name = c("Luke","Mark", "Hannah")R is case sensitive
R is case sensitive, which means that R makes a distinction between uppercase and lowercase letters for variables' names (e.g., 'score' and 'Score' are considered different names)
Example
> score <- c(8.5,7.5,9.2)
> score
[1] 8.5 7.5 9.2
> Score
Error: object 'Score' not found
Be careful when assigning variables' names and avoid confusion (!)
Accessing variables
You can retrieve or extract specific values or subsets from any data structure. The different data structures in R have distinct methods for accessing elements.
Vector [ ]
> score
[1] 8.5 7.5 9.2 6.3 8.7 7.2
> # second element
> score[2]
[1] 7.5
> # 1st and 3rd element
> score[c(1,3)]
[1] 8.5 9.2
Matrix [ , ]
> example_matrix
[,1] [,2] [,3]
[1,] 1.00 2.00 3.00
[2,] 1.33 2.33 3.33
[3,] 1.66 2.66 3.66
> # selecting element 1st row, 3rd col
> example_matrix[1,3]
[1] 3.00
> # selecting whole 1st row
> example_matrix[1,]
[1] 1.00 2.00 3.00
> # selecting whole 3rd column
> example_matrix[,3]
[1] 3.00 3.33 3.66
Operators : and -c( )
You can select contiguous elements of a vector or a matrix using the operator ':'. You can exclude elements (therefore selecting the remaining) by specifying the indices to exclude to the vector '-c( )'
Vector [ ]
> score
[1] 8.5 7.5 9.2 6.3 8.7 7.2
> # 1st, 2nd and 3rd element
> score[c(1:3)]
[1] 8.5 7.5 9.2
> # another way (excluding 4th,
> # 5th and 6th element)
> score[-c(4:6)]
[1] 8.5 7.5 9.2
Matrix [ , ]
> example_matrix
[,1] [,2] [,3]
[1,] 1.00 2.00 3.00
[2,] 1.33 2.33 3.33
[3,] 1.66 2.66 3.66
> # selecting first two rows and columns
> example_matrix[c(1,2),c(1,2)]
[,1] [,2]
[1,] 1.00 2.00
[2,] 1.33 2.33
> # another way
> # excluding 3rd row and 3rd column
> example_matrix[-3,-3]
[,1] [,2]
[1,] 1.00 2.00
[2,] 1.33 2.33
Accessing List and Data frame
[[ ]] or $
List
> sample_list <- list(name = name,
age = age,
score = score)
> sample_list
$name
[1] "Luke" "Mark" "Hannah"
$age
[1] 22 30 29
$score
[1] 8.5 7.5 9.2
# accessing with $ and object name
> sample_list$name
[1] "Luke" "Mark" "Hannah"
# accessing with [[ ]] and object name
> sample_list[["name"]]
[1] "Luke" "Mark" "Hannah"
# accessing with [[ ]] and index
> sample_list[[1]]
[1] "Luke" "Mark" "Hannah"
Data frame
> sample_df <- data.frame(name = name,
age = age,
score = score)
> sample_df
name age score
1 Luke 22 8.5
2 Mark 30 7.5
3 Hannah 29 9.2
# accessing with $ and column name
> sample_df$name
[1] "Luke" "Mark" "Hannah"
# accessing with [[ ]] and column name
> sample_df[["name"]]
[1] "Luke" "Mark" "Hannah"
# accessing with [[ ]] and index
> sample_df[[1]]
[1] "Luke" "Mark" "Hannah"
Calculating with variables
Variables can be used for performing any calculation that is possible given the data structure and the data type they are assigned to.
score
[1] 8.5 7.5 9.2
addition
> score + 1
[1] 9.5 8.5 10.2
subtraction
> score - 1
[1] 7.5 6.5 8.2
multiplication
> score * 2
[1] 17.0 15.0 18.4
division
> score / 10
[1] 0.85 0.75 0.92
sum( )
> sum(score)
[1] 25.2
mean( )
> mean(score)
[1] 8.4
max( )
> max(score)
[1] 9.2
min( )
> min(score)
[1] 7.5
score
[1] 8.5 7.5 9.2
addition
> score + 1
[1] 9.5 8.5 10.2
subtraction
> score - 1
[1] 7.5 6.5 8.2
multiplication
> score * 2
[1] 17.0 15.0 18.4
division
> score / 10
[1] 0.85 0.75 0.92
sum( )
> sum(score)
[1] 25.2
mean( )
> mean(score)
[1] 8.4
max( )
> max(score)
[1] 9.2
min( )
> min(score)
[1] 7.5
Function
Defining a function
A function is a reusable block of code that executes one or more specific tasks. A function is designed to take input values, called arguments, process them, and return an output.
> my_function_name <- function(arg1, arg2,
arg3, ...) {
# Body of the function with
# code to be exectuted
# ...
return(output) # return output object
}
- '
my_function_name' is the name of the function
- '
arg1', 'arg2' and 'arg3' are the function's arguments
- the body of the function contains the code that is executed
- '
return( )' is the statement used to specify the objects to be returned by the function
my_function_name' is the name of the functionarg1', 'arg2' and 'arg3' are the function's argumentsreturn( )' is the statement used to specify the objects to be returned by the functionExample with mean( )
The function 'mean( )' in R calculates the arithmetic mean (average) of a numeric vector
> # Define a numeric vector named 'numbers'
> numbers <- c(1,2,3,4,5,6,7,8,9,10)
> # calculate mean using 'mean( )' and assign it to 'average'
> average <- mean(x = numbers)
> # Print out 'average' variable
> average
[1] 5.5
Custom mean function
We can write our own function to compute the average of a numeric vector.
my_mean( )
> # Define custom function 'my_mean'
> my_mean <- function(x) {
total <- sum(x) # sum of elements of x
n <- length(x) # number of elements inside vector x
output <- total / n # mean stored in (local) object called 'output'
return(output) # return output object
}
# Using the custom function on the numeric vector 'numbers'
my_average <- my_mean(numbers)
> my_average
[1] 5.5
Sharing and reusing functions:
R packages
Installing and loading R packages
You can install a package from the official repository of R packages (CRAN) using the function install.packages("package_name"). You can load the library of functions from an installed package by using the function library(package_name)
Example with 'ggplot2' package
> # install package 'ggplot2' used for data visualization
> # (we will need it in the second part of the workshop!)
> install.packages("ggplot2")
> # NOTE:
> # at the beginning of the installation you may be asked to select
> # a mirror from which to download the package (56 is Netherlands)
> # you can choose whatever number
> # load library 'ggplot2' with library() function
> library(ggplot2)
Using functions from R packages
After loading the library with the function library(package_name), you can call the functions inside package_name:
(1) by using their name, name_function( )
(2) or with name_package : : name_function( )
> # loading library 'ggplot2'
> library(ggplot2)
> # (1) calling function 'ggplot( )' from 'ggplot2' package
> plot1 <- ggplot(data = data)
> # (2) calling function 'ggplot( )' using the syntax
> # name_package::name_function( )
> plot1 <- ggplot2::ggplot(data = data)
The syntax with : : is usually convenient when two or more loaded packages have functions with the same name
Help documentation
The help documentation refers to the detailed information available for functions, packages and datasets. It provides an essential support for users to understand the usage, functionality and parameters of functions of the installed packages. You can access the help documentation in R using the function help( ) or the operator ' ? '
help( ) and ?
> # help documentation of package "base"
> help(package = "ggplot2")
> # help documentation of function "ggplot( )" inside package "ggplot2"
> help(topic = "ggplot", package = "gglpot2")
> # or
> ?ggplot2::ggplot
> # when the library "ggplot2" is already loaded on the workspace
> ?ggplot
Warnings and Errors
Warnings and errors are messages printed on the console to inform users about potential issues during the execution of a chunk of code or a function.
A warning message refer to potential problems with the execution of a command but the execution can still proceed without being interrupted.
> # Warning about squared root of a negative number
> sqrt(-5)
[1] NaN
Warning message:
In sqrt(-5) : NaNs produced
An error message refers to a critical issue that prevent the command from being executed. Something went wrong and the operation cannot be successfully completed.
> # Error about a variable that has not been defined
> result <- x - 5
Error: object 'x' not found
R Workspace Management
Workspace
The term workspace refers to the working environment where R objects and data are stored during an R session. It includes all the objects that are currently loaded and available for use. You can use the command 'ls( )' to print on console the variable names' associated to all the loaded variables in the workspace.
> ls()
[1] "age" "data_sample" "example_matrix" "name"
[5] "sample_df" "sample_list" "score"
Working directory
The working directory refers to the location in your computer where R by default looks for (reads) and saves (writes) files.
getwd( )
> getwd()
[1] "/Users/giuseppe"
setwd( )
> setwd("/Users/giuseppe/Desktop/")
> getwd()
[1] "/Users/giuseppe/Desktop"
Importing and exporting tabular data
Importing tabular data
You can read text files (.txt) and CSV files (Comma-Separated Value, .csv) using the functions read.table( ) or read.csv( )
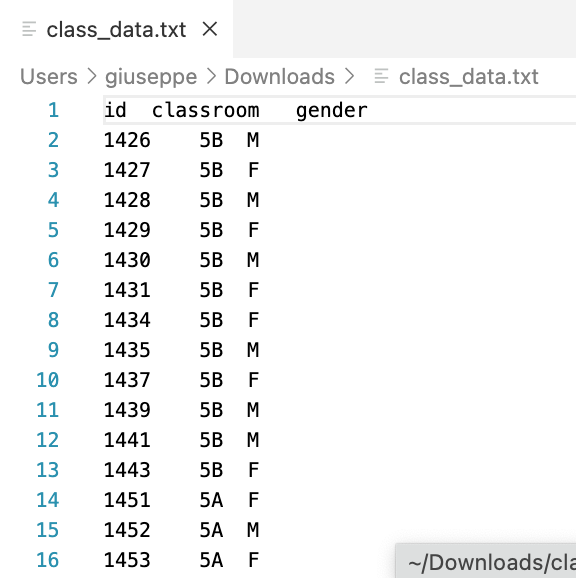
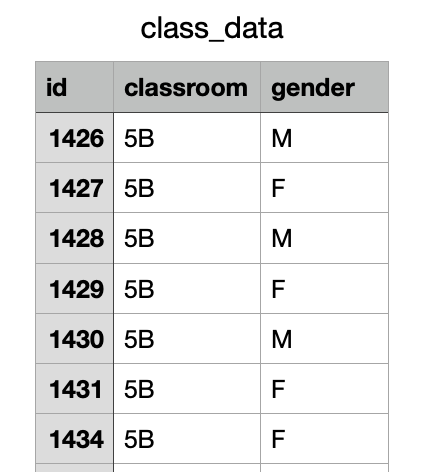
.txt file
> class_df <- read.table(
file = "/Users/giuseppe/Downloads/class_data.txt",
header = TRUE,
sep = "",
dec = "."
)
# head( ) prints out the first 6 elements of
# any data structure
head(class_df)
id classroom gender
1 1426 5B M
2 1427 5B F
3 1428 5B M
4 1429 5B F
5 1430 5B M
6 1431 5B F
.csv file
> class_df <- read.csv(
file = "/Users/giuseppe/Downloads/class_data.csv",
header = TRUE,
sep = ",",
dec = "."
)
head(class_df)
id classroom gender
1 1426 5B M
2 1427 5B F
3 1428 5B M
4 1429 5B F
5 1430 5B M
6 1431 5B F
Other useful functions for importing tabular data in R:
| Data to import | R function | package |
|---|---|---|
| .csv files that use a comma as decimal point (dec = ' , ') and a semicolon as field separator (sep = ' ; ') | read.csv2( ) | utils |
| Excel worksheets from an Excel workbook | read_excel( ) | readxl |
| read.xlsx( ) | xlsx | |
| SPSS datasets | read_sav( ) | haven |
Exporting tabular data
You can save tabular format into text files (.txt) or CSV files using the functions write.table( ) or write.csv( )
write.table( )
> head(class_df)
id classroom gender
1 1426 5B M
2 1427 5B F
3 1428 5B M
4 1429 5B F
5 1430 5B M
6 1431 5B F
> write.table(x = class_df,
file = "class_df.txt",
dec = ".",
sep = "")
Other functions for exporting tabular data:
| Type of data | R function | package |
|---|---|---|
| .csv files with comma as decimal points semicolon as field separator | write.csv2( ) | utils |
| Excel worksheet | write.xlsx( ) | xlsx |
| SPSS datasets | write_sav( ) | haven |
Summary of
Part I
Intro to R language and console (RStudio panels)
Objects: data structures and functions
Creating and accessing variables
R packages, help documentation, and workspace
Importing and exporting tabular data
Intro to R language and console (RStudio panels)
Objects: data structures and functions
Creating and accessing variables
R packages, help documentation, and workspace
Importing and exporting tabular data