Introduction to R
(Part II)
Giuseppe Arena
Tilburg University
2024-01-18
Welcome back!
Goals of part II
![]()
![]()
exploring
data
![]()
visualizing
data
![]()
running
statistical analyses


exploring
data
visualizing
data
running
statistical analyses
Data exploration
Loading data from R package into workspace
We use the dataset HolzingerSwineford1939 available inside the package lavaan.
> # install 'lavaan' package
> install.packages("lavaan", dependencies = TRUE)
>
> # loading 'lavaan' package
> library(lavaan)
This is lavaan 0.6-17
lavaan is FREE software! Please report any bugs.
>
> # load dataset in the workspace, you can check this with the function 'ls()'
> data("HolzingerSwineford1939")
>
> # use a shorter symbolic name for the dataset
> dataset1 <- HolzingerSwineford1939
>
> # re-assign the column 'sex' as a factor
> dataset1$sex <- factor(dataset1$sex)
Description of the dataset
The Holzinger and Swineford dataset (1939) consists of mental ability test scores of seventh- and eighth-grade children from two different schools (Pasteur and Grant-White). For the complete documentation use ?HolzingerSwineford1939.
| Variable name | Description |
|---|---|
| id | Identifier |
| sex | Gender |
| ageyr | Age, year part |
| agemo | Age, month part |
| school | School (Pasteur or Grant-White) |
| grade | Grade |
| Variable name | Description | Factor |
|---|---|---|
| x1 | Visual perception | |
| x2 | Cubes | visual |
| x3 | Lozenges | |
| x4 | Paragraph comprehension | |
| x5 | Sentence completion | textual |
| x6 | Word meaning | |
| x7 | Speeded addition | |
| x8 | Sepeeded counting of dots | speed |
| x9 | Speeded discrimination straight and curved capitals | |
Basic functions for a data.frame object
Basic functions for a data.frame object
dim( ), colnames( ), rownames( )
> # dimensions of the dataset
> dim(dataset1) # 301 rows (observations) and 15 columns (variables)
[1] 301 15
>
> # column names of 'dataset1'
> colnames(dataset1) # 'names(dataset1)' returns the same information
[1] "id" "sex" "ageyr" "agemo" "school" "grade" "x1" "x2" [...]
>
> # row names of 'dataset1'
> rownames(dataset1)
[1] "1" "2" "3" "4" "5" "6" "7" "8" "9" "10" [...]
head( )
> # first six rows of a data.frame
> head(dataset1[,c(1:6)])
id sex ageyr agemo school grade
1 1 1 13 1 Pasteur 7
2 2 2 13 7 Pasteur 7
3 3 2 13 1 Pasteur 7
4 4 1 13 2 Pasteur 7
5 5 2 12 2 Pasteur 7
6 6 2 14 1 Pasteur 7
tail( )
> # last six rows of a data.frame
> tail(dataset1[,c(1:6)])
id sex ageyr agemo school grade
296 345 1 13 3 Grant-White 8
297 346 1 13 5 Grant-White 8
298 347 2 14 10 Grant-White 8
299 348 2 14 3 Grant-White 8
300 349 1 14 2 Grant-White 8
301 351 1 13 5 Grant-White NA
Structure and summary of a data.frame
str( ) and summary( )
> # structure of the data.frame: dimensions, name and data type of each column in the data.frame
> str(dataset1[,c(1:6)])
'data.frame': 301 obs. of 6 variables:
$ id : int 1 2 3 4 5 6 7 8 9 11 ...
$ sex : Factor w/ 2 levels "1","2": 1 2 2 1 2 2 1 2 2 2 ...
$ ageyr : int 13 13 13 13 12 14 12 12 13 12 ...
$ agemo : int 1 7 1 2 2 1 1 2 0 5 ...
$ school: Factor w/ 2 levels "Grant-White",..: 2 2 2 2 2 2 2 2 2 2 ...
$ grade : int 7 7 7 7 7 7 7 7 7 7 ...
>
> # summary of the data.frame object (prints out a summary per each columns of the data.frame)
> ## for quantitative variables: Min - 1st Quartile - Median - Mean - 3rd Quartile - Max - NA's
> ## for qualitative (categorical) variables: frequency values
> summary(dataset1[,c(1:6)])
id sex ageyr agemo school grade
Min. : 1.0 1:146 Min. :11 Min. : 0.000 Grant-White:145 Min. :7.000
1st Qu.: 82.0 2:155 1st Qu.:12 1st Qu.: 2.000 Pasteur :156 1st Qu.:7.000
Median :163.0 Median :13 Median : 5.000 Median :7.000
Mean :176.6 Mean :13 Mean : 5.375 Mean :7.477
3rd Qu.:272.0 3rd Qu.:14 3rd Qu.: 8.000 3rd Qu.:8.000
Max. :351.0 Max. :16 Max. :11.000 Max. :8.000
NA's :1
Filtering rows of a data.frame with subset( )
Filtering rows of a data.frame with subset( )
Example: subset of students from school "Pasteur", selecting only information about age (ageyr), and visual variables (x1, x2, x3).
> # Filtering a dataset
> subset1 <- subset(x = dataset1,
+ subset = school == "Pasteur",
+ select = c(ageyr, x1, x2, x3))
>
> head(subset1)
ageyr x1 x2 x3
1 13 3.333333 7.75 0.375
2 13 5.333333 5.25 2.125
3 13 4.500000 5.25 1.875
4 13 5.333333 7.75 3.000
5 12 4.833333 4.75 0.875
6 14 5.333333 5.00 2.250
The logical operator == (double equal sign) is used for checking equality between two objects. Instead, to check whether two objects are not equal, use the operator !=.
Filtering with more conditions:
logical operators & (AND), | (OR)
Filtering with more conditions
Example 1: subset of students of the 7th grade AND from school "Pasteur", selecting only information about sex (sex), and visual variables (x1, x2, x3).
> subset2 <- subset(x = dataset1,
+ subset = grade == 7 & school == "Pasteur",
+ select = c(sex, x1, x2, x3))
> head(subset2)
sex x1 x2 x3
1 1 3.333333 7.75 0.375
2 2 5.333333 5.25 2.125
3 2 4.500000 5.25 1.875
4 1 5.333333 7.75 3.000
5 2 4.833333 4.75 0.875
6 2 5.333333 5.00 2.250
The logical operator & (AND) compares two logical values (the one on the left and the one on the right of the symbol) and returns a TRUE only if both logical values are TRUE.
Filtering with more conditions
Example 2: subset of students from the school Pasteur AND with age less than 13 OR greater than 14 years, selecting sex (sex), age (ageyr), and visual variables (x1, x2, x3).
> subset3 <- subset(x = dataset1,
+ subset = (school == "Pasteur") & (ageyr < 13 | ageyr > 14),
+ select = c(sex,ageyr, x1, x2, x3))
> head(subset3)
sex ageyr x1 x2 x3
5 2 12 4.833333 4.75 0.875
7 1 12 2.833333 6.00 1.000
8 2 12 5.666667 6.25 1.875
10 2 12 3.500000 5.25 0.750
11 1 12 3.666667 5.75 2.000
12 1 12 5.833333 6.00 2.875
The logical operator | (OR) compares two logical values (the one on the left and the one on the right of the symbol) and returns a TRUE if at least one of the two logical values is TRUE.
colMeans( )
colMeans( ) allows to calculate the mean for each column of a data.frame (or matrix).
> # Function 'colMeans( )'
>
> # subset school == "Pasteur"
> subset_pasteur <- subset(x = dataset1,
+ subset = school == "Pasteur",
+ select = c(ageyr, x1, x2, x3) )
> # subset school == "Grant-White"
> subset_grantwhite <- subset(x = dataset1,
+ subset = school == "Grant-White",
+ select = c(ageyr, x1, x2, x3) )
>
> # colMeans
> colMeans(subset_pasteur, na.rm = TRUE)
ageyr x1 x2 x3
13.250000 4.941239 5.983974 2.487179
> colMeans(subset_grantwhite, na.rm = TRUE)
ageyr x1 x2 x3
12.724138 4.929885 6.200000 1.995690
Another similar function is colSums( ), which returns the sum value per each column in the data.frame (or matrix).
apply( )
apply(X, MARGIN, FUN) allows to apply the function FUN to the MARGIN of the object X (a data.frame or a matrix)
> # Applying a function (FUN) to columns/rows (MARGIN) of a data.frame/matrix (X)
>
> # using function 'apply( )' with arguments
> # X, the data
> # MARGIN, 1 for applying FUN by row, 2 for applying FUN by columns
> # FUN, the function to apply to each row or column (e.g., sd(), function to
> # calculate standard deviation)
>
> apply(X = subset_pasteur,
+ MARGIN = 2,
+ FUN = sd,
+ na.rm = TRUE)
ageyr x1 x2 x3
1.063318 1.185003 1.230224 1.163818
>
> apply(X = subset_grantwhite,
+ MARGIN = 2,
+ FUN = sd,
+ na.rm = TRUE)
ageyr x1 x2 x3
0.9681222 1.1523039 1.1112587 1.0396244
tapply( )
tapply(X, INDEX, FUN) allows to apply a function FUN on a variable X per each level of another variable INDEX (categorical).
> # mean of 'ageyr' per each level of variable 'school'
> tapply(X = dataset1$ageyr, INDEX = dataset1$school, FUN = mean)
Grant-White Pasteur
12.72414 13.25000
>
> # standard deviation of 'x1', 'x2', and 'x3' per each level of variable 'school'
> tapply(X = dataset1$x1, INDEX = dataset1$school, FUN = sd, na.rm = TRUE)
Grant-White Pasteur
1.152304 1.185003
> tapply(X = dataset1$x2, INDEX = dataset1$school, FUN = sd, na.rm = TRUE)
Grant-White Pasteur
1.111259 1.230224
> tapply(X = dataset1$x3, INDEX = dataset1$school, FUN = sd, na.rm = TRUE)
Grant-White Pasteur
1.039624 1.163818
Frequency tables
The function table( ) in R is used to create tables returning the frequency (counts) of the levels of one or more categorical variables. The frequency table can be created on the levels of one categorical variable (one-way table), on two categorical variables (two-way table) or on more categorical variables. The function prop.table( ) calculates the relative frequencies from a frequency table..
table( )
> # Contingency tables - table( )
>
> # (1) One-way contingency table of
> # 'school'
> table(dataset1$school)
Grant-White Pasteur
145 156
> # 145 students from "Grant-White",
> # 156 from "Pasteur"
>
> # (2) Two-way contingency table of
> # 'school' and 'sex'
> table(dataset1$school, dataset1$sex)
1 2
Grant-White 72 73
Pasteur 74 82
prop.table( )
> # Relative frequency table - prop.table( )
> # (1) for one-way tables
> school_tab <- table(dataset1$school)
> prop.table(x = school_tab)
Grant-White Pasteur
0.4817276 0.5182724
>
> # (2) for two-way tables
> school_sex_tab <- table(dataset1$school,
+ dataset1$sex)
> prop.table(x = school_sex_tab, margin = 1)
1 2
Grant-White 0.4965517 0.5034483
Pasteur 0.4743590 0.5256410
Note: for one-way tables (1), the relative frequency will be calculated using the total of the table (145/301 = 0.4817 and 156/301 = 0.5183). For two-way tables (2), the user can specify which 'margin' (= 1 by rows or, = 2 by columns) to refer for the total (e.g., (2) with margin = 1).
Variance and covariance
var( )
> # (1) var(x) to calculate variance
> # of vector 'ageyr'
> var(x = dataset1$ageyr)
[1] 1.103322
>
> # (2) var(x,y) to calculate covariance
> # beween variables 'ageyr' and 'x1'
> var(x = dataset1$ageyr, y = dataset1$x1)
[1] -0.07354743
cov( )
> # (1) cov(x,y) to calculate
> # covariance between 'ageyr' and 'x1'
> cov(x = dataset1$ageyr, y = dataset1$x1)
[1] -0.07354743
>
> # (2) cov(x) calculates covariance matrix
> # between 'ageyr' and 'x1'
> # (supplying a data.frame to x)
> cov(x = dataset1[,c("ageyr","x1")])
ageyr x1
ageyr 1.10332226 -0.07354743
x1 -0.07354743 1.36289774
The standard deviation of a variable can be calculated using the function sd( ).
Correlation
cov2cor( )
> # calculate covariance matrix with cov(x)
> cov_matrix <- cov(x = dataset1[,c("ageyr",
+ "x1")])
>
> # convert to correlation matrix
> # V is the input covariance matrix
> cov2cor(V = cov_matrix)
cor( )
> # (1) cor(x,y) calculates the correlation
> # between 'ageyr' and 'x1'
> cor(x = dataset1$ageyr, y = dataset1$x1)
[1] -0.05997699
>
> # (2) cor(x) correlation matrix of the
> # variables in a data.frame
> cor(x = dataset1[,c("ageyr","x1","x2","x3")])
ageyr x1 x2 x3
ageyr 1.000000 -0.059977 -0.01526 0.03718
x1 -0.059977 1.000000 0.29735 0.44067
x2 -0.015260 0.297346 1.00000 0.33985
x3 0.037179 0.440668 0.33985 1.0000
Data visualization
R packages: 'graphics' and 'ggplot2'
R packages: 'graphics' and 'ggplot2'
'graphics'
> # plot function
> plot(x, # data
+ y, # data
+ type, # points (p), lines (l), etc..
+ main, # plot title
+ xlab, # x-axis label
+ ylab, # y-axis label
+ pch, # point style
+ lwd, # line width
+ lty, # line style
+ cex, # point size
+ col # color,
+ ...) # etc.
Input arguments avialable also for other plotting functions,
depending on the type of plot
'ggplot2'
> # plot function with 'ggplot2'
> # concatenation of ggplot objects by '+'
> ggplot(data, # a data.frame
+ mapping = aes(x,y)) +
+ # mapping defines an aes() object
+ # geom_point( ) add points to
+ # the ggplot object
+ geom_point(color, size) +
+ labs(title,
+ x,
+ y) +
+ theme_bw() # theme that defines the
+ # aspect of the plot
Other functions of the type geom_xzy() allow to display different types of plots
Scatter plot
The scatter plot is used for visualizing the relationship between two numeric variables. Individual data points are plotted on a two-dimensional graph. Each data point is represented by a dot, and its position on the graph depends on the values of the two variables used in the plot. With a scatter plot, it is possible to detect outliers and identify potential patterns.
plot(x, y, type = "p")
> # Scatter plot
> plot(x = dataset1$x1,
+ y = dataset1$x2,
+ type = "p", # 'p' = 'points'
+ main = "Scatter plot 'x1' vs. 'x2'",
+ xlab = "x1",
+ ylab = "x2",
+ pch = 1,
+ cex = 0.5,
+ col = "gray50")

geom_point( )
> # Scatter plot with 'ggplot2'
> ggplot(data = dataset1,
+ mapping = aes(x = x1, y = x2)) +
+ geom_point(color = "gray50", size = 0.8) +
+ labs(title = "Scatter plot 'x1' vs. 'x2'",
+ x = "x1",
+ y = "x2") +
+ theme_bw()

plot(x, y, type = "p")
> # Scatter plot
> plot(x = dataset1$x1,
+ y = dataset1$x2,
+ type = "p", # 'p' = 'points'
+ main = "Scatter plot 'x1' vs. 'x2'",
+ xlab = "x1",
+ ylab = "x2",
+ pch = 1,
+ cex = 0.5,
+ col = "gray50")

geom_point( )
> # Scatter plot with 'ggplot2'
> ggplot(data = dataset1,
+ mapping = aes(x = x1, y = x2)) +
+ geom_point(color = "gray50", size = 0.8) +
+ labs(title = "Scatter plot 'x1' vs. 'x2'",
+ x = "x1",
+ y = "x2") +
+ theme_bw()

Line plot
The line plot is used for visualizing data points connected by straight lines. The line shows the trend of a variable over an ordered variable, for instance the time. A line plot is commonly used for time series data, where the x-axis represents the time variable, and the y-axis represents the variable of interest.
plot(x, y, type = "l" or "b")
> # Line plot
> # some random data
> time <- 1:20
> y <- 3.5 + time*0.4 + rnorm(n=length(time))
> plot(x = time,
+ y = y,
+ type = "l", # 'l' = 'lines'
+ main = "Line plot of 'y'",
+ xlab = "time",
+ ylab = "y",
+ col = "gray50")

geom_line( )
> # Line plot with 'ggplot2'
> ggplot(data = data.frame(time = time, y = y),
+ mapping = aes(x = time, y = y)) +
+ geom_line(color = "gray50") +
+ labs(title = "Line plot of 'y'",
+ x = "time",
+ y = "y") +
+ theme_bw()

plot(x, y, type = "l" or "b")
> # Line plot
> # some random data
> time <- 1:20
> y <- 3.5 + time*0.4 + rnorm(n=length(time))
> plot(x = time,
+ y = y,
+ type = "l", # 'l' = 'lines'
+ main = "Line plot of 'y'",
+ xlab = "time",
+ ylab = "y",
+ col = "gray50")

geom_line( )
> # Line plot with 'ggplot2'
> ggplot(data = data.frame(time = time, y = y),
+ mapping = aes(x = time, y = y)) +
+ geom_line(color = "gray50") +
+ labs(title = "Line plot of 'y'",
+ x = "time",
+ y = "y") +
+ theme_bw()

Bar plot
The bar plot is used to visualize categorical variables (e.g., 'school'). The observed frequencies of the levels of a categorical variable are described by vertical bars, whose height corresponds to the frequency value of each level of the variable. The bars are arranged along one axis (usually the x-axis, i.e. the horizontal axis).
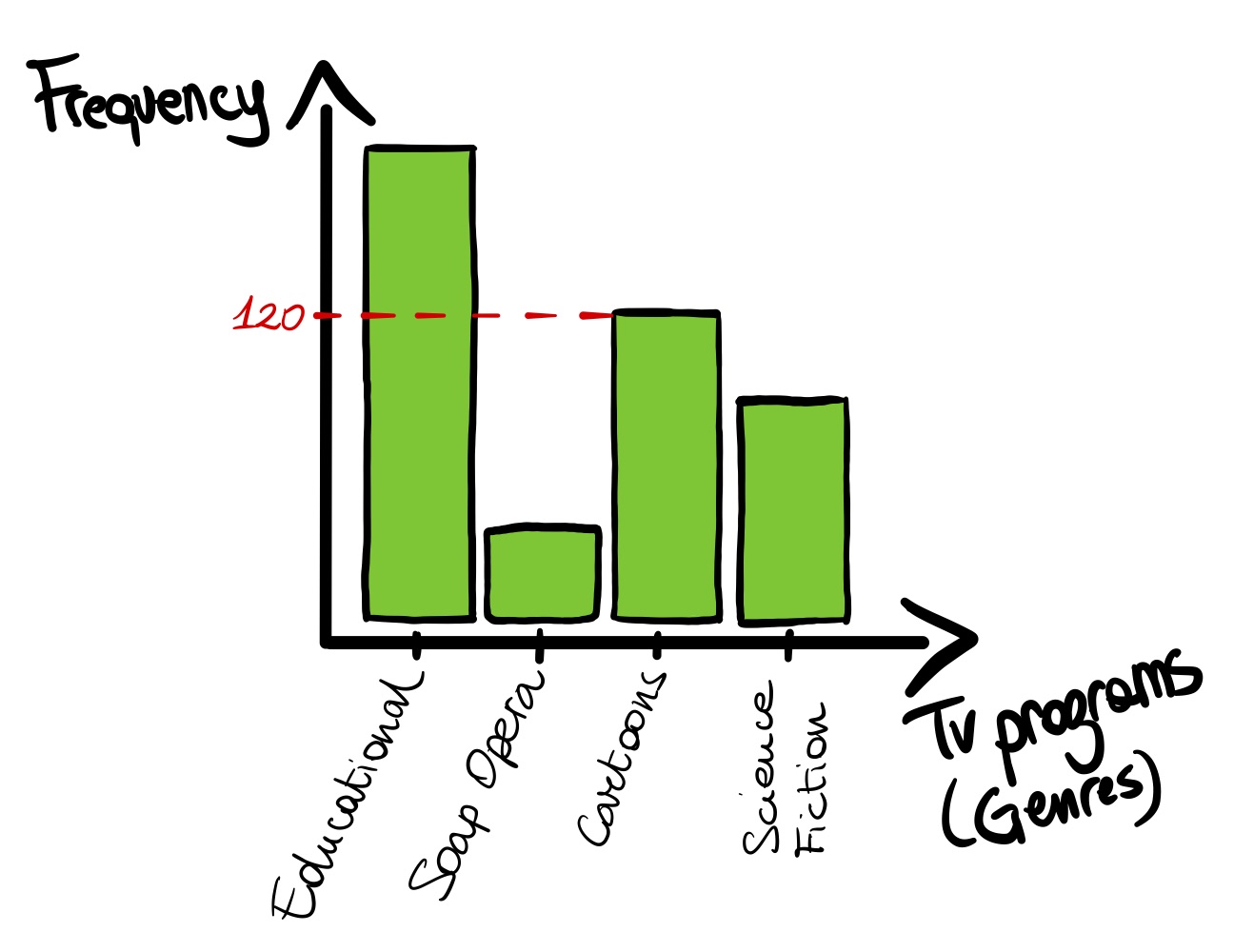
barplot( )
> # Bar plot
> frequency_school <- table(dataset1$school)
> barplot(height = frequency_school,
+ horiz = FALSE,
+ main = "Bar plot of 'school'",
+ xlab = "school",
+ ylab = "number of students",
+ ylim = c(0,
+ max(frequency_school+10)),
+ col = "aquamarine3")
> box() # adds a square around the plot

geom_bar( )
> # Bar plot with 'ggplot2'
> ggplot(data = dataset1,
+ mapping = aes(x = school)) +
+ geom_bar(stat = "count",
+ fill = "aquamarine3") +
+ labs(title = "Bar plot of 'school'",
+ x = "school",
+ y = "number of students") +
+ scale_x_continuous(breaks=c(11:16)) +
+ theme_bw()

barplot( )
> # Bar plot
> frequency_school <- table(dataset1$school)
> barplot(height = frequency_school,
+ horiz = FALSE,
+ main = "Bar plot of 'school'",
+ xlab = "school",
+ ylab = "number of students",
+ ylim = c(0,
+ max(frequency_school+10)),
+ col = "aquamarine3")
> box() # adds a square around the plot

geom_bar( )
> # Bar plot with 'ggplot2'
> ggplot(data = dataset1,
+ mapping = aes(x = school)) +
+ geom_bar(stat = "count",
+ fill = "aquamarine3") +
+ labs(title = "Bar plot of 'school'",
+ x = "school",
+ y = "number of students") +
+ scale_x_continuous(breaks=c(11:16)) +
+ theme_bw()

Histogram
The histogram is used to visualize the distribution of the values of a numeric variable. It divides the range of the variable into intervals, also called bins, and counts the number of observations that fall into each interval. The resulting histogram consists of a sequence of adjacent rectangles whose heights correspond to the frequency (or density) of observations falling into a specific interval.
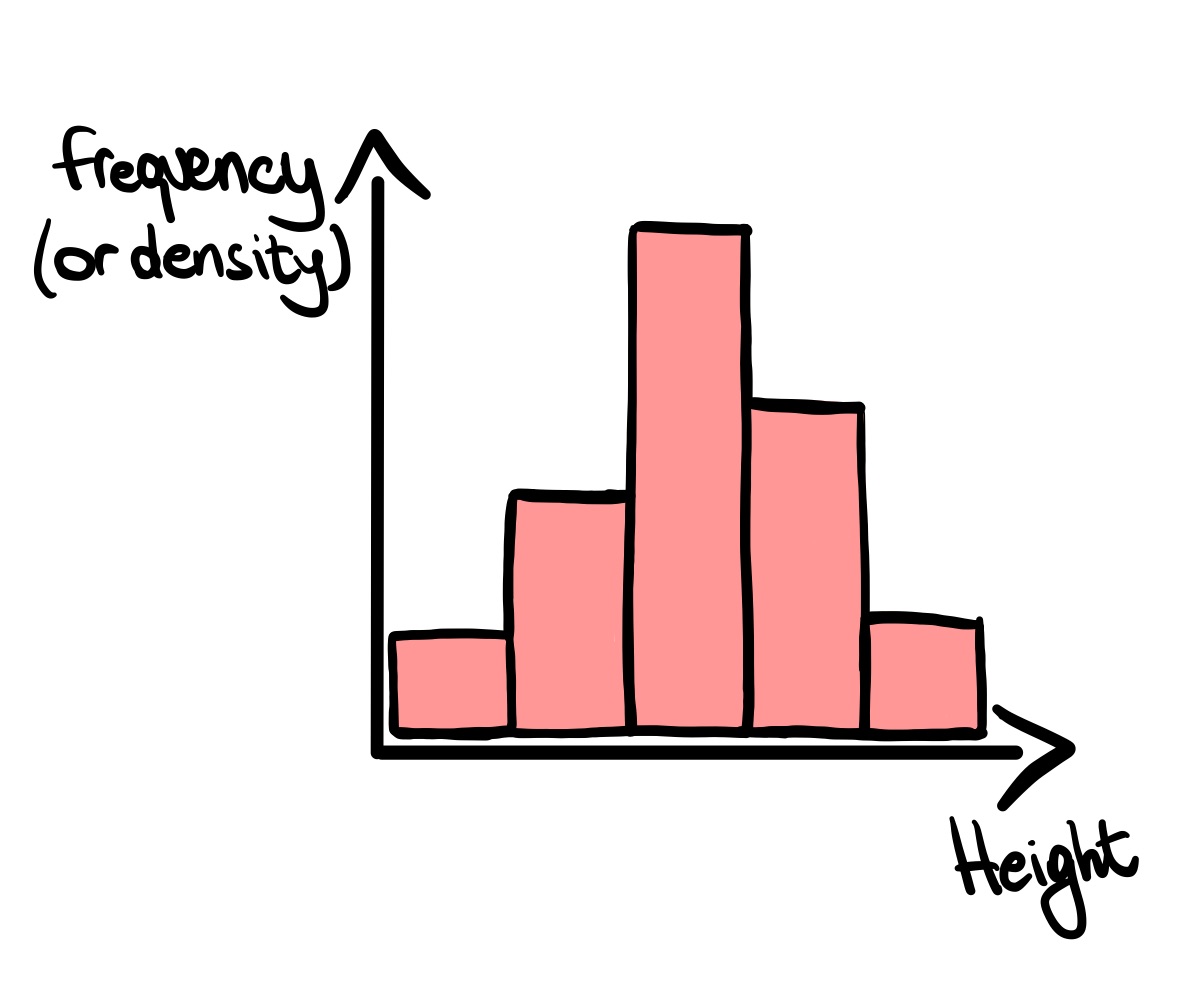
hist( )
> # Histogram of 'x1' ('visual perception')
> hist(x = dataset1$x1,
+ freq = TRUE,
+ main = "Visual perception",
+ xlab = "value",
+ ylab = "frequency",
+ col = "lavender")
> box()

geom_histogram( )
> # Histogram with 'ggplot2'
> ggplot(data = dataset1,
+ mapping = aes(x = x1)) +
+ geom_histogram(bins = 9,
+ fill = "lavender",
+ color = "black") +
+ labs(title = "Visual perception",
+ x = "value",
+ y = "frequency") +
+ theme_bw()

hist( )
> # Histogram of 'x1' ('visual perception')
> hist(x = dataset1$x1,
+ freq = TRUE,
+ main = "Visual perception",
+ xlab = "value",
+ ylab = "frequency",
+ col = "lavender")
> box()

geom_histogram( )
> # Histogram with 'ggplot2'
> ggplot(data = dataset1,
+ mapping = aes(x = x1)) +
+ geom_histogram(bins = 9,
+ fill = "lavender",
+ color = "black") +
+ labs(title = "Visual perception",
+ x = "value",
+ y = "frequency") +
+ theme_bw()

Density plot
Similar to a histogram, the density plot represents the distribution of a continuous variable. A density plot is a smooth curve which that estimates probability density function of a continuous variable.
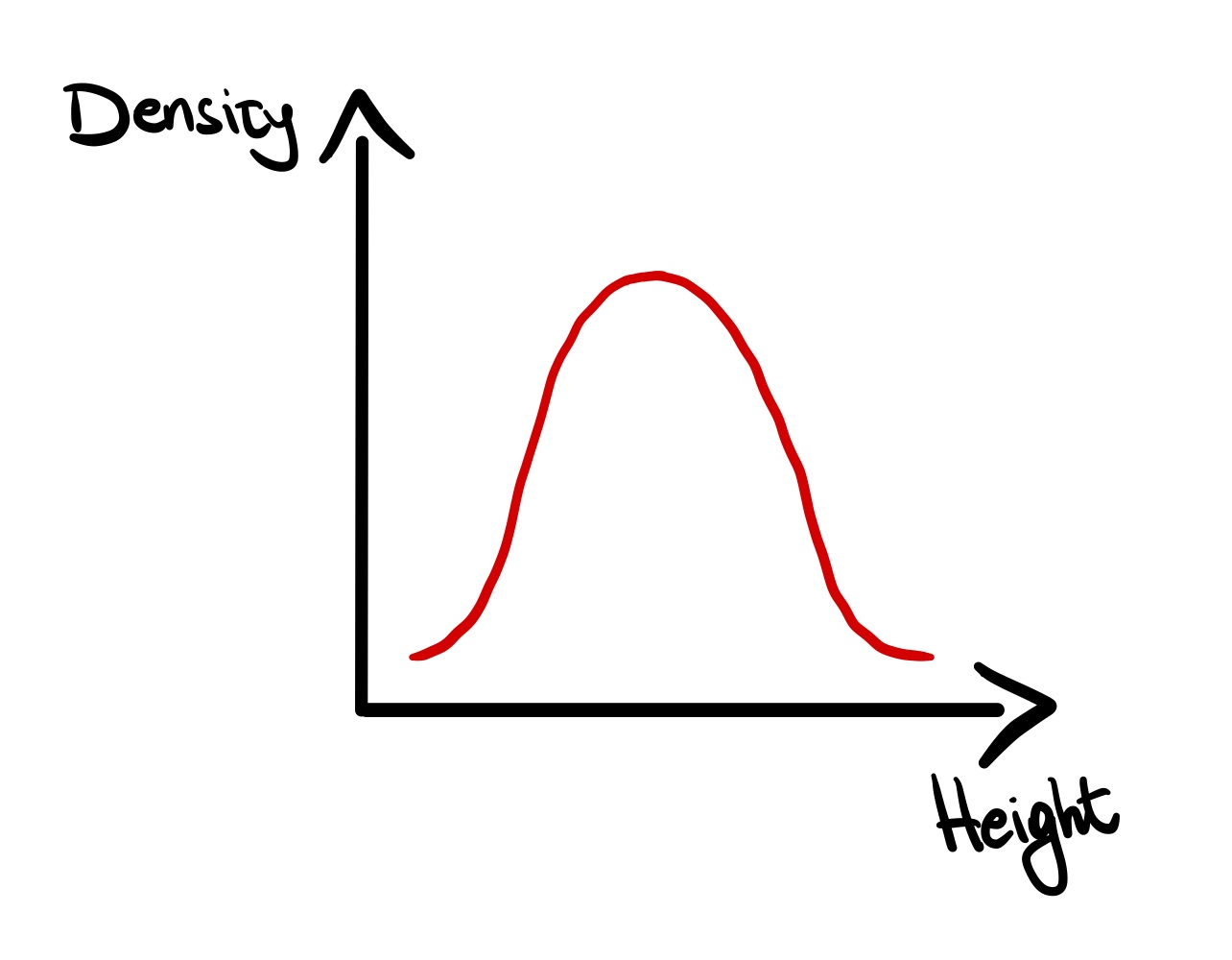
plot( x = density( ) )
> # Density plot of 'x1'
> plot(x = density(dataset1$x1),
+ main = "Visual perception",
+ xlab = "value",
+ ylab = "density",
+ lwd = 2)

geom_density( )
> # Density with 'ggplot2'
> ggplot(data = dataset1,
+ mapping = aes(x = x1)) +
+ geom_density(color = "black",
+ lwd = 1) +
+ labs(title = "Visual perception",
+ x = "value",
+ y = "density") +
+ theme_bw()

plot( x = density( ) )
> # Density plot of 'x1'
> plot(x = density(dataset1$x1),
+ main = "Visual perception",
+ xlab = "value",
+ ylab = "density",
+ lwd = 2)

geom_density( )
> # Density with 'ggplot2'
> ggplot(data = dataset1,
+ mapping = aes(x = x1)) +
+ geom_density(color = "black",
+ lwd = 1) +
+ labs(title = "Visual perception",
+ x = "value",
+ y = "density") +
+ theme_bw()

Box plot
A boxplot displays the distribution of numeric variables, their central tendency and variability. It provides a visual summary of the distribution by including interquartile range (length of the box), median (line inside the box), whiskers (the minimum and maximum values from the quartiles), and outliers (all points outside the whiskers). The boxplot is useful for outliers identification and for comparing the distribution of a variable across different levels (groups) of a categorical variable.
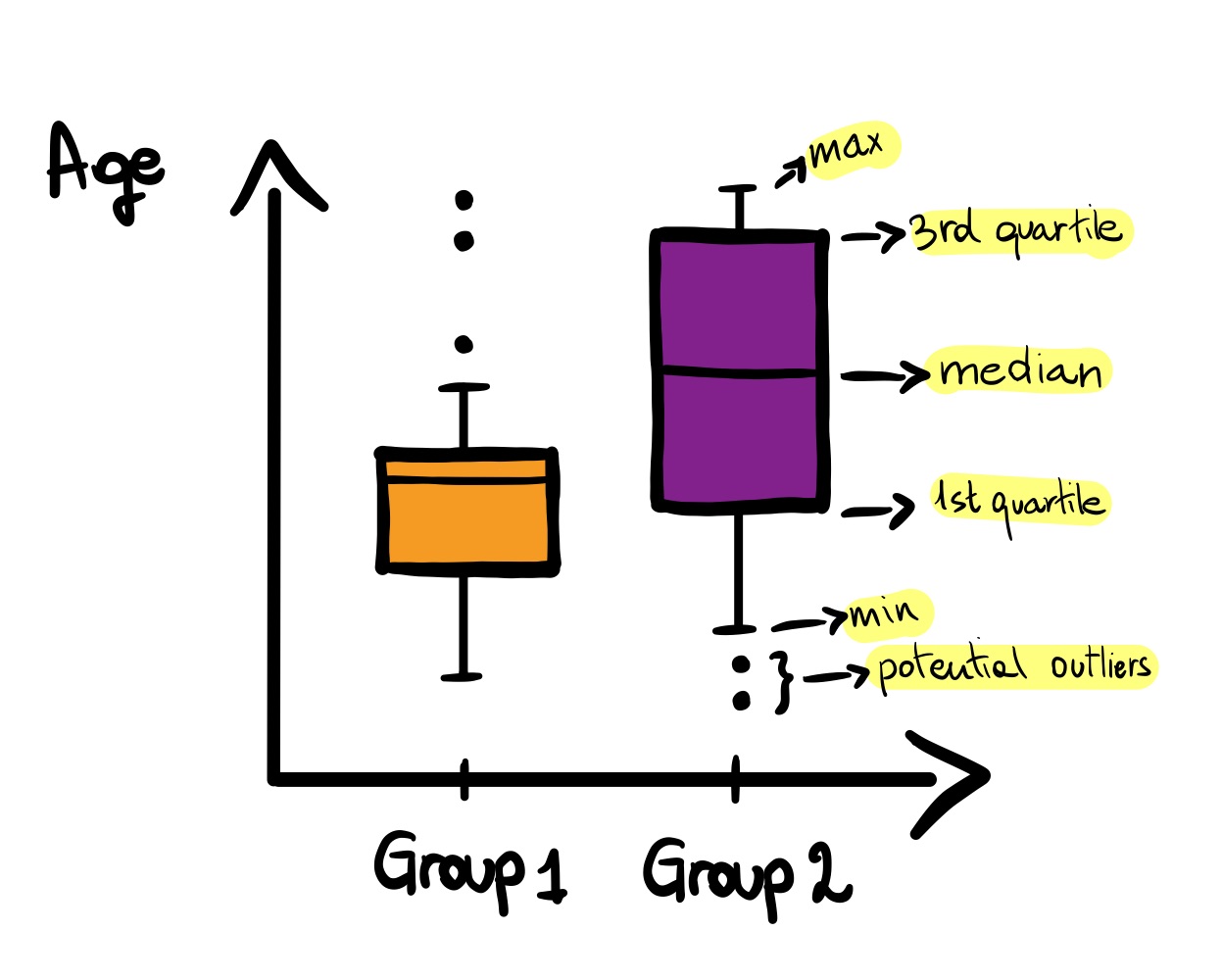
boxplot('x' or 'formula' )
> # Box plot
> # boxplot(x = , ...) # one variable only
> boxplot(
+ formula = dataset1$ageyr ~ dataset1$school,
+ main = "Age (years)",
+ xlab = "school",
+ ylab = "ageyr",
+ col = c("orange","turquoise4"),
+ horizontal = FALSE)
>
> legend("top",
+ legend = levels(dataset1$school),
+ fill = c("orange","turquoise4"))

geom_boxplot( )
> # Box plot with 'ggplot2'
> ggplot(data = dataset1,
+ mapping = aes(x = school,
+ y = ageyr)) +
+ geom_boxplot(mapping = aes(fill = school),
+ color = "black") +
+ labs(title = "Age (years)",
+ x = "school",
+ y = "ageyr") +
+ scale_fill_manual(values = c("orange",
+ "turquoise4")) +
+ theme_bw()

boxplot('x' or 'formula' )
> # Box plot
> # boxplot(x = , ...) # one variable only
> boxplot(
+ formula = dataset1$ageyr ~ dataset1$school,
+ main = "Age (years)",
+ xlab = "school",
+ ylab = "ageyr",
+ col = c("orange","turquoise4"),
+ horizontal = FALSE)
>
> legend("top",
+ legend = levels(dataset1$school),
+ fill = c("orange","turquoise4"))

geom_boxplot( )
> # Box plot with 'ggplot2'
> ggplot(data = dataset1,
+ mapping = aes(x = school,
+ y = ageyr)) +
+ geom_boxplot(mapping = aes(fill = school),
+ color = "black") +
+ labs(title = "Age (years)",
+ x = "school",
+ y = "ageyr") +
+ scale_fill_manual(values = c("orange",
+ "turquoise4")) +
+ theme_bw()

geom_violin( ) with 'ggplot2'
> # Violin plot with 'ggplot2'
> ggplot(data = dataset1,
+ mapping = aes(x = school,
+ y = x1)) +
+ geom_violin(mapping = aes(fill = school),
+ color = "black") +
+ labs(title = "Visual perception (x1)",
+ x = "school",
+ y = "Visual perception (x1)") +
+ geom_boxplot(width=0.1) +
+ scale_fill_manual(values = c("orange",
+ "turquoise4"))+
+ theme_bw()

> # Violin plot with 'ggplot2'
> ggplot(data = dataset1,
+ mapping = aes(x = school,
+ y = x1)) +
+ geom_violin(mapping = aes(fill = school),
+ color = "black") +
+ labs(title = "Visual perception (x1)",
+ x = "school",
+ y = "Visual perception (x1)") +
+ geom_boxplot(width=0.1) +
+ scale_fill_manual(values = c("orange",
+ "turquoise4"))+
+ theme_bw()

Correlation plot
A correlation plot shows the pairwise correlation coefficients between the variables of a dataset, which helps with understanding the relationships between multiple variables. The R package corrplot provides a function to create correlation plots.
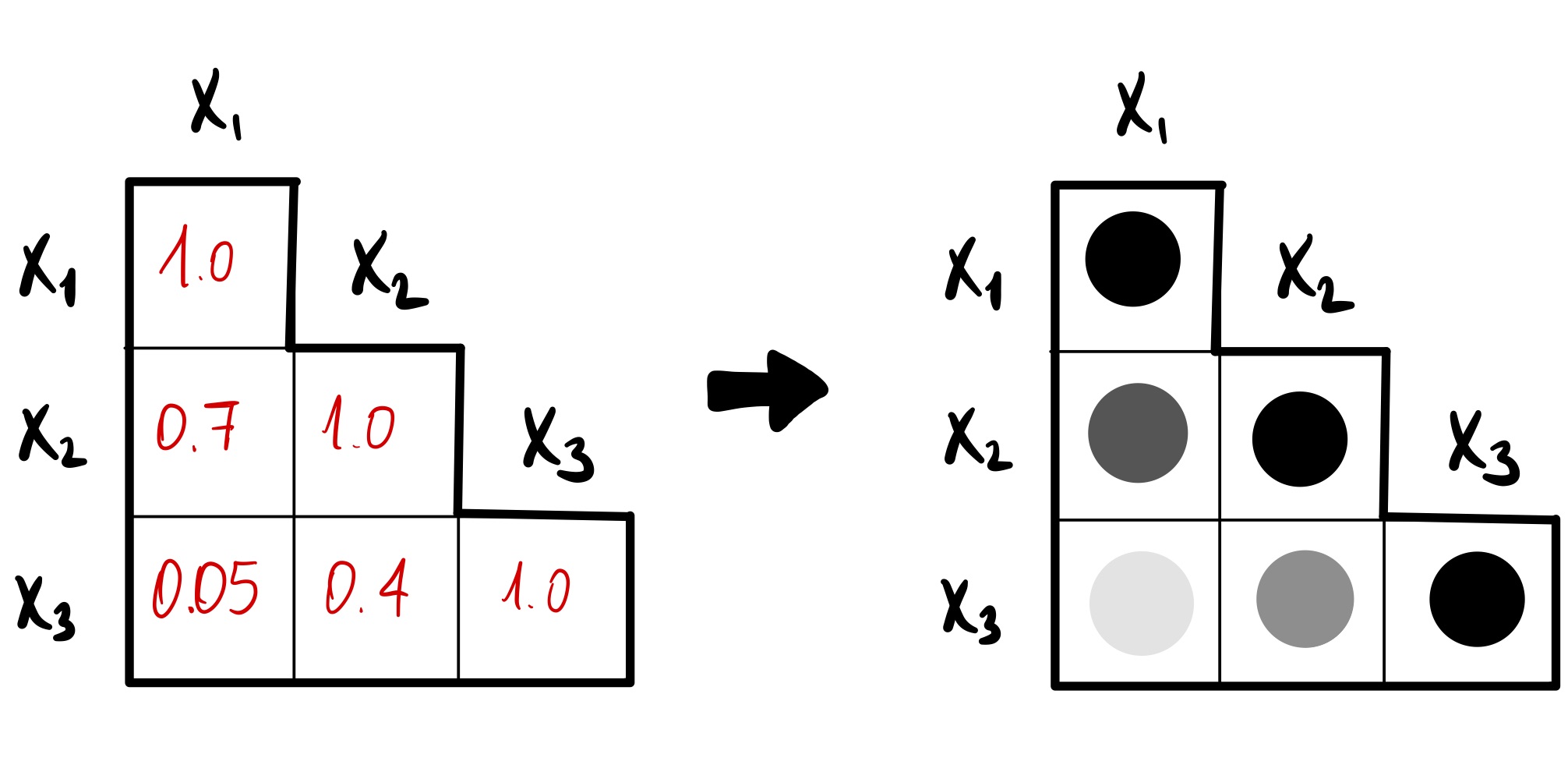
corrplot( ) with 'corrplot'
> # Correlation plot
>
> # Install and load 'corrplot' package
> install.packages("corrplot")
> library(corrplot)
>
> # correlation matrix of x1, x2, ..., x9
> correlation_mat <- cor(dataset1[,c(7:15)])
>
> # Correlation plot with 'corrplot'
>
> # (1) method = "circle"
> corrplot(correlation_mat,
+ method = "circle",
+ type="lower")
>
> # (2) method = "number"
> corrplot(correlation_mat,
+ method = "number",
+ type="lower")


> # Correlation plot
>
> # Install and load 'corrplot' package
> install.packages("corrplot")
> library(corrplot)
>
> # correlation matrix of x1, x2, ..., x9
> correlation_mat <- cor(dataset1[,c(7:15)])
>
> # Correlation plot with 'corrplot'
>
> # (1) method = "circle"
> corrplot(correlation_mat,
+ method = "circle",
+ type="lower")
>
> # (2) method = "number"
> corrplot(correlation_mat,
+ method = "number",
+ type="lower")


Theme (ggplot2)
In ggplot2, themes control several visual aspects of a plot, for instance, axis labels, the title, the legend, the background color, grid lines and other characteristics. The user can apply an available theme or a customized one. Some of the available themes are theme_classic( ), theme_bw( ), and theme_minimal( ). Themes are helpful for customizing a plot's styling and matching a scientific journal's publication standards. The function theme( ) allows for the customization of different aspects of the plot.
Applying a custom theme
> # Theme (ggplot2)
> # (1) create ggplot object 'boxplot_x1'
> boxplot_x1 <- ggplot(data = dataset1,
+ mapping = aes(x = school,
+ y = x1)) +
+ geom_boxplot(mapping = aes(fill = school),
+ color = "black") +
+ labs(title = "Visual perception",
+ x = "school",
+ y = "Visual perception") +
+ scale_fill_manual(values = c("orange",
+ "turquoise4"))
>
> # (2) create ggplot theme 'theme_custom_1'
> theme_custom_1 <- theme_bw() +
+ theme(axis.title = element_text(size = 20,
+ vjust = 0.5),
+ line = element_line(linetype = "dashed"),
+ plot.title = element_text(size = 40,
+ hjust = 0.5),
+ legend.position = "bottom",
+ legend.direction = "horizontal",
+ legend.text = element_text(size = 15),
+ legend.title = element_text(size = 15))
> # (3) add theme to the plot
> boxplot_x1 + theme_custom_1
Saving plots ...
... with RStudio
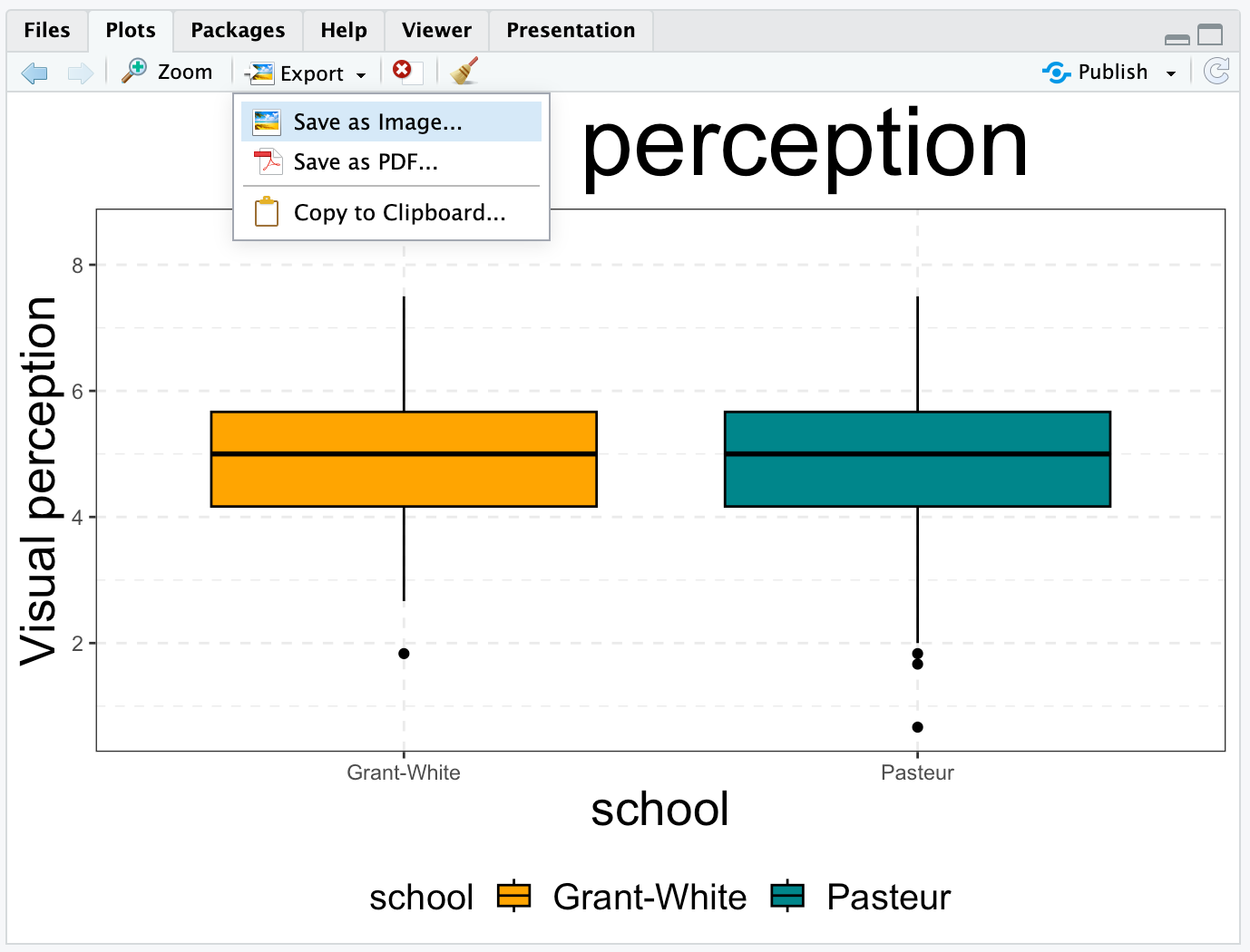
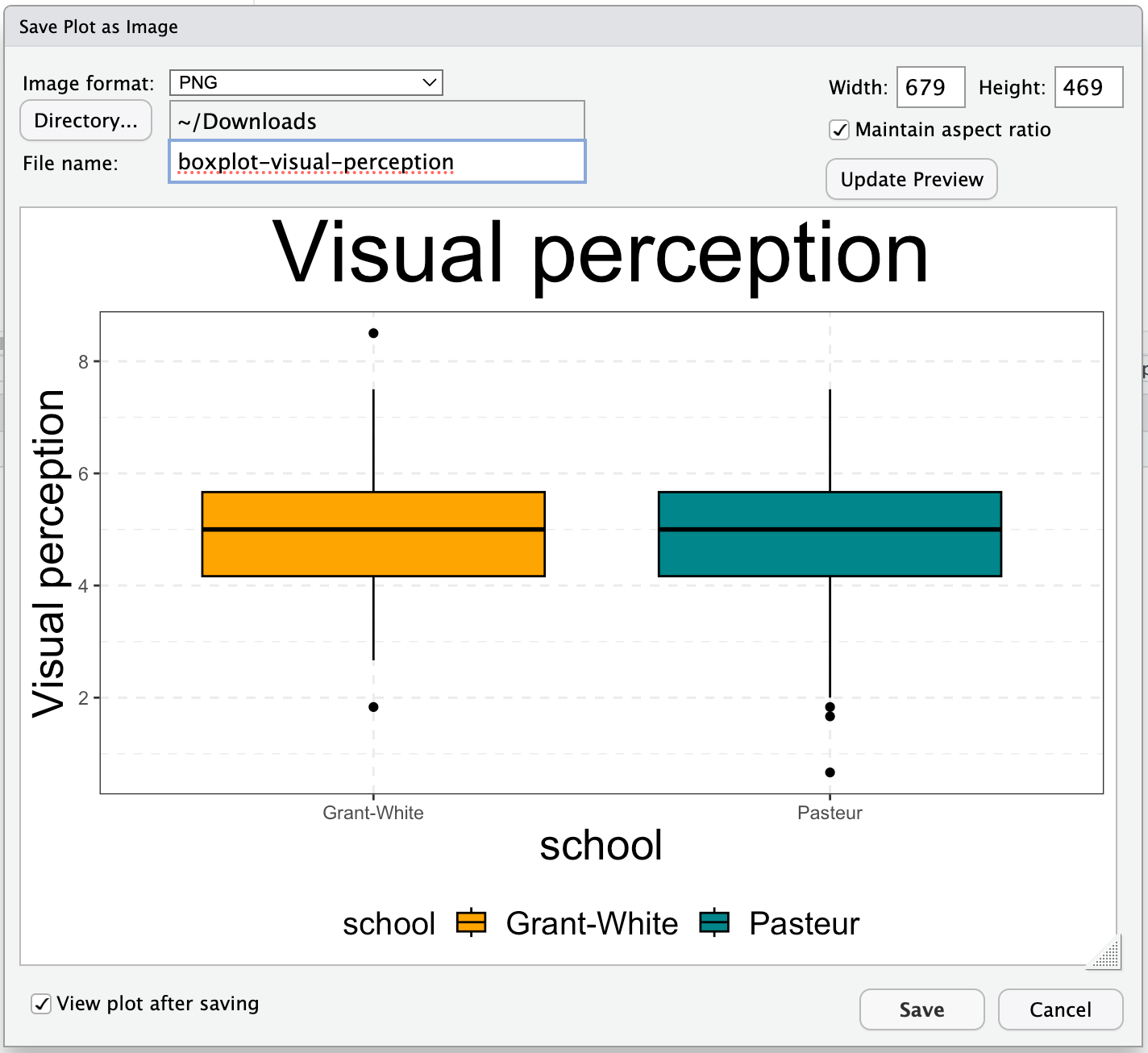
Saving plots ...
... from R console
using grDevices
> # method (2): with 'png( )'
> # saving image into current working directory
> png(filename = "visual-perception.png",
+ width = 600,
+ height = 400,
+ units = "px")
> print(boxplot_x1 + theme_custom_1)
> dev.off() # closing R device for graphics
Other extensions are available, using the functions jpeg( ), bmp( ), tiff( ), svg( ), and pdf( )
using ggplot2
> # method (1): with 'ggsave( )'
> ggsave(filename = "visual-perception.png",
+ width = 600,
+ height = 400,
+ units = "px")
Other extensions are available, such as .JPEG, .BMP, .TIFF, .SVG, and .PDF
Statistical Analyses in R
Linear Regression

> # Run a simple linear regression
> lm_model <- lm(formula = x6 ~ x4,
> data = dataset1)
> # Inspect the analysis results.
> summary(lm_model)
Call:
lm(formula = x6 ~ x4, data = dataset1)
Residuals:
Min 1Q Median 3Q Max
-1.88636 -0.52249 -0.01577 0.46656 3.07582
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept)0.15613 0.12647 1.234 0.218
x4 0.66302 0.03863 17.164 <2e-16 ***
---
Signif. codes: 0 ‘***’ 0.001 ‘**’
0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
Residual standard error: 0.779 on 299 degrees
of freedom
Multiple R-squared: 0.4963,
Adjusted R-squared: 0.4946
F-statistic: 294.6 on 1 and 299 DF,
p-value: < 2.2e-16
Linear Regression

> # Visualize the regression line
> ggplot(data = dataset1,
+ mapping = aes(x = x4,
+ y = x6)) +
+ geom_point(color = "gray50") +
+ geom_smooth(method = "lm",
+ formula = y ~ x,
+ se = FALSE,
+ color = "red") +
+ theme_bw() +
+ labs(title = "Scatter plot and regression line",
+ x = "Paragraph comprehension (x4)",
+ y = "Word meaning (x6)")
Diagnostics plot
> # Model diagnostics
> # par() to set grid 2x2 with mfrow()
> par(mfrow=c(2,2))
>
> # plot of 'lm' object
> plot(lm_model,
+ cex = 0.8, # size of the points
+ pch = 19, # point style
+ col = rgb(red = 127,
+ green = 127,
+ blue = 127,
+ alpha = 90,
+ maxColorValue = 255), # color
+ lwd = 2) # line width

Multiple Linear Regression
> # Run a multiple linear regression
> lm_multiple <- lm(formula = x1 ~ x2 + x3 + x4,
> data = dataset1)
>
> # Inspect the analysis results.
> summary(lm_multiple)
Call:
lm(formula = x1 ~ x2 + x3 + x4, data = dataset1)
Residuals:
Min 1Q Median 3Q Max
-3.6957 -0.6529 0.1000 0.6616 2.3413
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 2.40878 0.31567 7.631 3.20e-13 ***
x2 0.13247 0.05133 2.581 0.0103 *
x3 0.35936 0.05349 6.718 9.38e-11 ***
x4 0.29789 0.04946 6.023 5.04e-09 ***
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
Residual standard error: 0.979 on 297 degrees of freedom
Multiple R-squared: 0.3039, Adjusted R-squared: 0.2968
F-statistic: 43.21 on 3 and 297 DF, p-value: < 2.2e-16
Structural Equation Models (SEM)
> library(lavaan)
Multiple Linear Regression in SEM framework
Model specification and estimation
> # (1) specify model, same as with lm()
> model <- "x1 ~ 1 + x2 + x3 + x4"
> # (2) run estimation
> lm_sem_model <- sem(model = model,
+ data = dataset1)
Summary
> summary(lm_sem_model) # (3) inspect results
Estimator ML
Optimization method NLMINB
Number of model parameters 5
Number of observations 301
Model Test User Model:
Test statistic 0.000
Degrees of freedom 0
Parameter Estimates:
Standard errors Standard
Information Expected
Information saturated (h1) model Structured
Regressions:
Estimate Std.Err z-value P(>|z|)
x1 ~
x2 0.132 0.051 2.598 0.009
x3 0.359 0.053 6.763 0.000
x4 0.298 0.049 6.064 0.000
Intercepts:
Estimate Std.Err z-value P(>|z|)
.x1 2.409 0.314 7.682 0.000
Variances:
Estimate Std.Err z-value P(>|z|)
.x1 0.946 0.077 12.268 0.000
SEM framework with three latent variables
SEM framework with three latent variables
Model specification and estimation
> # (1) Specify SEM model with three latent variables.
> model_2 <- "
+ # Latent variables
+ latent1 = ~ x1 + x2 + x3
+ latent2 = ~ x4 + x5 + x6
+ latent3 = ~ x7 + x8 + x9
+
+ # Covariance between latent variables
+ latent1 ~~ latent2
+ latent1 ~~ latent3
+ latent2 ~~ latent3
+ "
> # (2) Run the analysis
> sem_model_2 <- sem(model = model_2, data = dataset1)
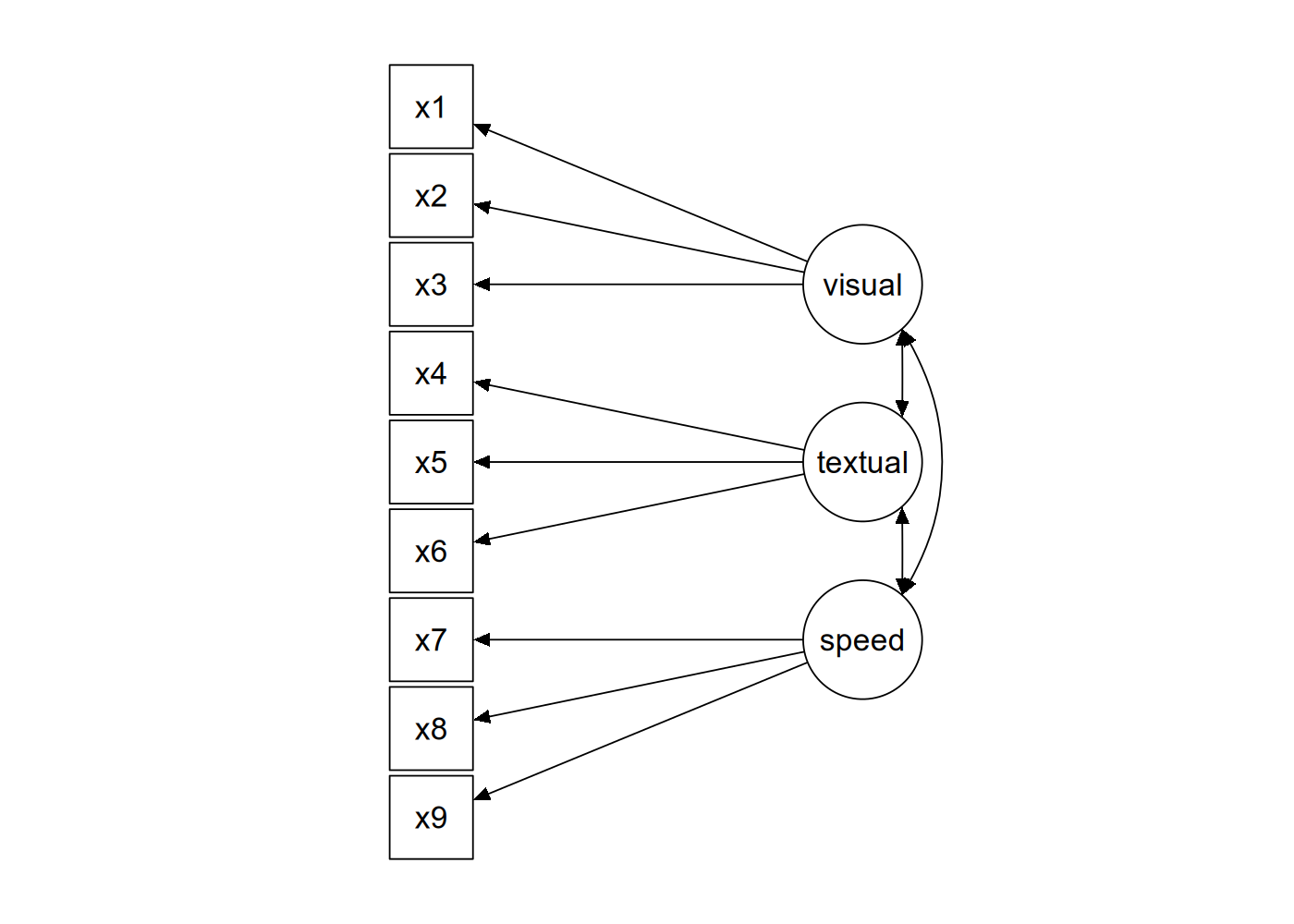
content inspired by tutorial at lavaan.ugent.be/tutorial/cfa.html
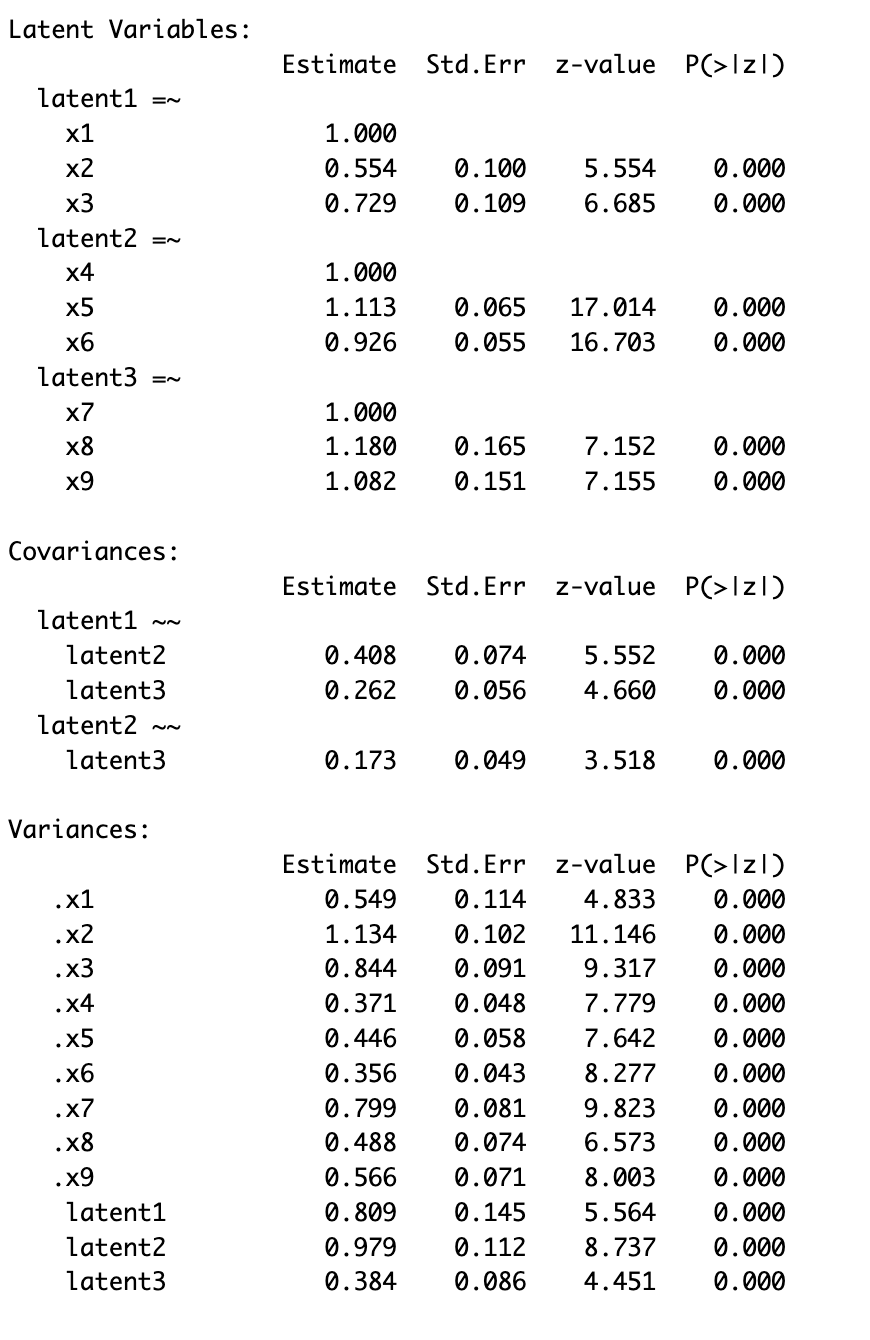
Summary SEM with latent variables
> # summary with fit measures
> summary(sem_model_2)
> round(
+ fitMeasures(sem_model_2)[c("chisq",
+ "df",
+ "pvalue",
+ "cfi",
+ "rmsea",
+ "srmr")], 3)
chisq df pvalue cfi rmsea srmr
85.306 24.000 0.000 0.931 0.092 0.065
+ # print out more fit measures with
+ # summary(sem_model_2, fit.measures = TRUE)
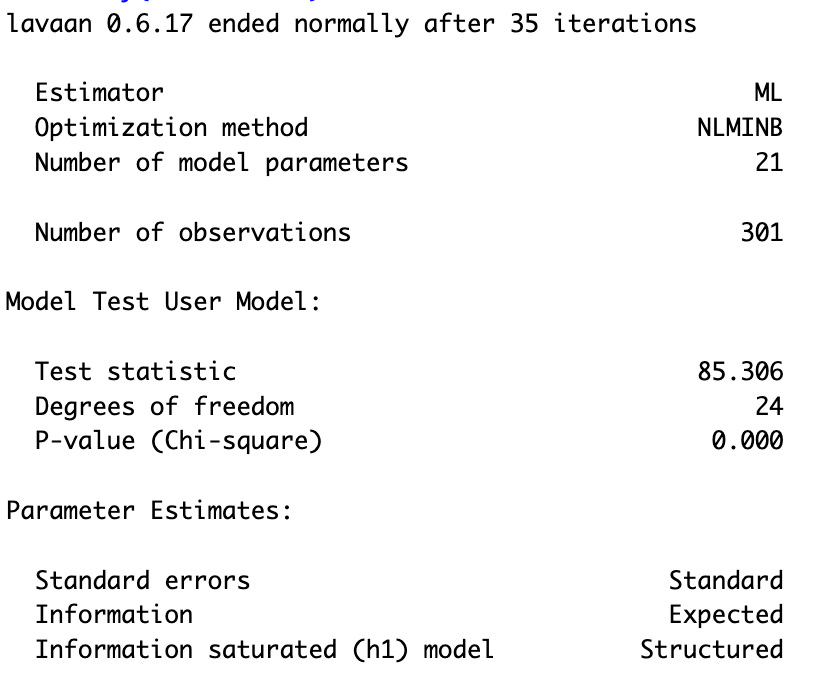

Visualize SEM model
> library(semPlot)
Visualize SEM model
> # Plot the SEM model with three latent variables
> semPaths(sem_model_2,
+ what = "path",
+ whatLabels = "est")

Linear Mixed-Effects Models
> library(lme4)
Random Intercept
Random intercept per school: model specification and estimation
> # Fit a linear mixed-effects model with
> # random intercepts for school
> lme_model <- x6 ~ 1 + (1|school) + sex +
+ ageyr + x4 + x5
> model <- lmer(formula = lme_model, data = dataset1)
> summary(model)
\[ \text{random intercept} \\ \text{per school:} \\ (1 | \text{school} )\]
Summary of the model
Linear mixed model fit by REML ['lmerMod']
Formula: x6 ~ 1 + (1 | school) + sex + ageyr + x4 + x5
Data: dataset1
REML criterion at convergence: 661.6
Scaled residuals:
Min 1Q Median 3Q Max
-2.2242 -0.6463 -0.0017 0.6048 3.4045
Random effects:
Groups Name Variance Std.Dev.
school (Intercept) 0.001672 0.04089
Residual 0.498582 0.70610
Number of obs: 301, groups: school, 2
Fixed effects:
Estimate Std. Error t value
(Intercept) -0.370401 0.582357 -0.636
sex2 -0.134552 0.083085 -1.619
ageyr -0.006922 0.040591 -0.171
x4 0.368365 0.051866 7.102
x5 0.365919 0.047045 7.778
Correlation of Fixed Effects:
(Intr) sex2 ageyr x4
sex2 -0.202
ageyr -0.965 0.152
x4 -0.044 -0.107 0.035
x5 -0.260 0.058 0.112 -0.719
Note: p-value's are not reported in the summary. The 'lmerTest' package runs with the function lmerTest : : lmer( ) (same name as in 'lme4' package) returns the p-value's (this is only available for Linear Mixed-Effects Models, not for Generalized Linear Mixed-Effects Models).
Diagnostics
> # Diagnostics of the model
>
> # Homoscedasticity of residuals
> plot(lme_model,
+ pch = 19,
+ cex = 0.8,
+ main = "Residuals",
+ )
>
> # normality - Q-Q plot
> qqnorm(residuals(lme_model))
> qqline(residuals(lme_model))


Mixed-Effects Model
Consider the variables: intercept, id (student), school, sex, x1, x2 and x3. Assume that for each student the abilities x1, x2, and x3 are measured multiple times.
| Effect | Model specification (formula) |
|---|---|
| random intercept per school | x1 ~ 1 + (1|school) |
| random intercept per school and sex | x1 ~ 1 + (1|school:sex) |
| fixed and random slope of x2 per student | x1 ~ x2 + (x2|id) |
| fixed and random slope of x2 and x3 per student | x1 ~ x2 + x3 + (x2 + x3|id) |
Summary of
Part II
Data exploration
Data visualization (with 'graphics' and 'ggplot2')
Structural Equation Models with 'lavaan' package
Linear Mixed-Effects models with 'lme4' package
Data exploration
Data visualization (with 'graphics' and 'ggplot2')
Structural Equation Models with 'lavaan' package
Linear Mixed-Effects models with 'lme4' package
Thanks for taking part in this workshop!
Happy R coding!

Resources
R coding:
Data visualization:
SEM with 'lavaan':
Linear Mixed-effects models:
R coding:
Data visualization:
SEM with 'lavaan':
Linear Mixed-effects models: